Clusters and Virtual Machines
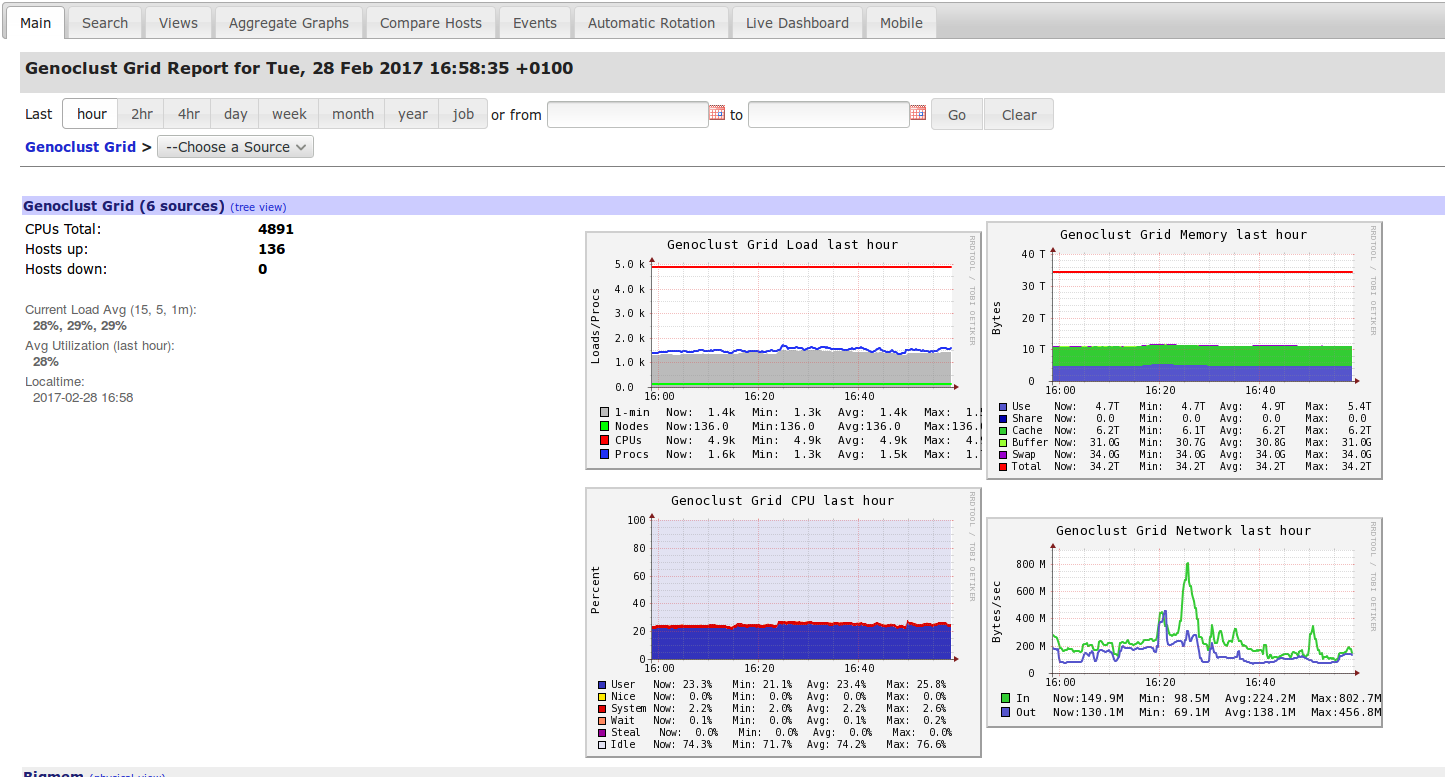
To see resources consumption (CPU, memory, …) of Genobioinfo cluster , you can use our Genobioinfo Ganglia system.
To see resources consumption (CPU, memory, …) of virtual machine, you can use our Genologin Ganglia system.
Some help to understand Ganglia view here.